georgebuilds/livewire-molecule
| Install | |
|---|---|
composer require georgebuilds/livewire-molecule |
|
| Latest Version: | v2.1.0 |
| PHP: | ^8.2 |
| License: | MIT |
| Last Updated: | May 5, 2026 |
| Links: | GitHub · Packagist |
Livewire Molecule
A Laravel Livewire component for 3D molecular visualization powered by 3DMol.js
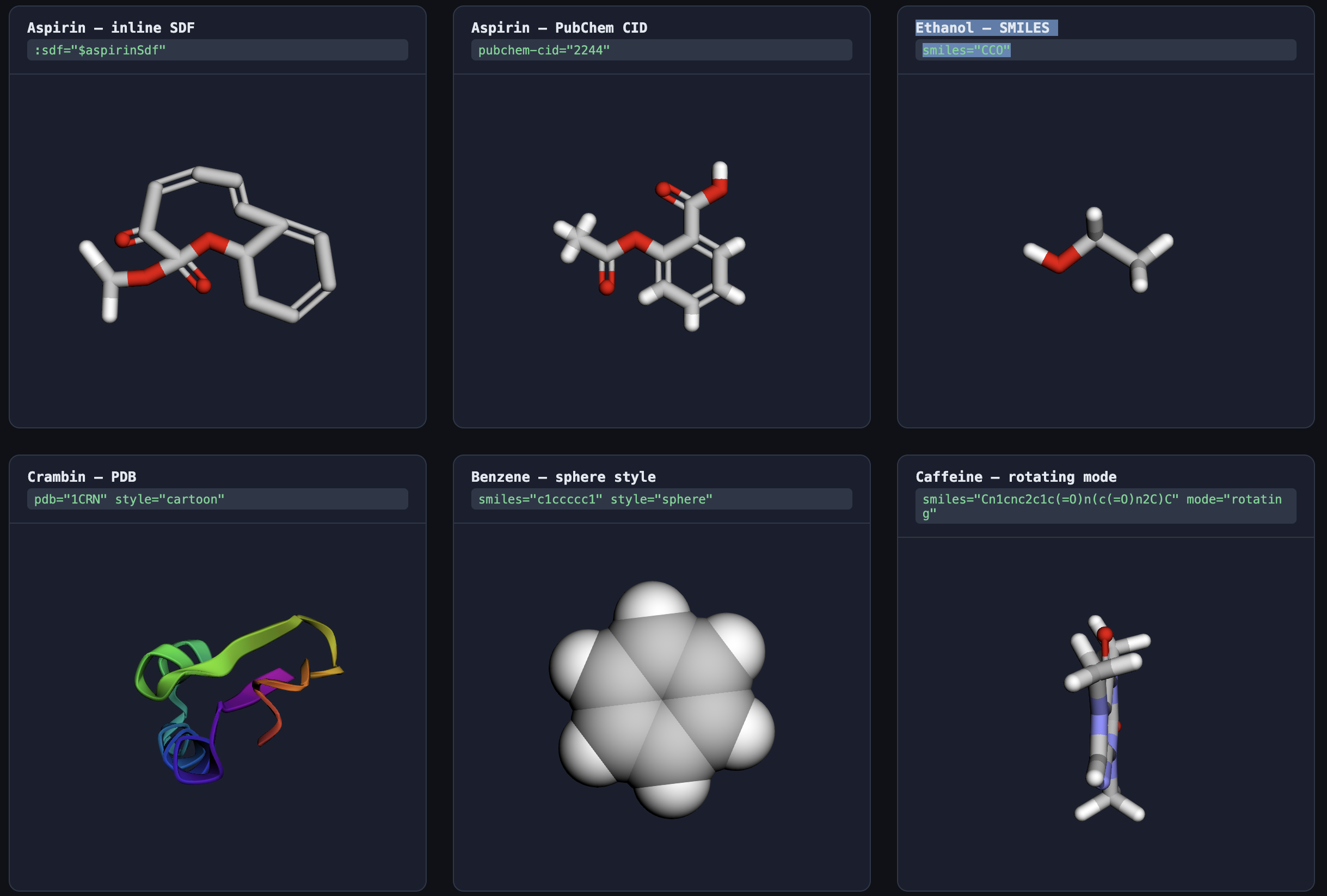
Features
- 🧪 Multiple input formats: SMILES, InChI, PDB ID, PubChem CID, or raw SDF data
- 🎨 Multiple visualization styles: stick, sphere, cartoon, line, ball-and-stick
- 🔄 Three display modes: interactive, rotating, static
- ⚡ Reactive updates with Livewire
- 🎯 Automatic 3D coordinate generation from SMILES/InChI
Requirements
- PHP 8.2+
- Laravel 11, 12, or 13
- Livewire 4.x (requires Laravel 11+)
- Alpine.js (included with Livewire)
Installation
composer require georgebuilds/livewire-molecule
Optionally publish the config file:
php artisan vendor:publish --tag=molecule-config
Usage
Basic Usage
{{-- From SMILES notation --}}
<livewire:molecule smiles="CCO" />
{{-- From PDB ID --}}
<livewire:molecule pdb="1CRN" />
{{-- From PubChem CID --}}
<livewire:molecule pubchem-cid="2244" />
{{-- From InChI --}}
<livewire:molecule inchi="InChI=1S/C2H6O/c1-2-3/h3H,2H2,1H3" />
{{-- From raw SDF data --}}
<livewire:molecule :sdf="$sdfContent" />
Display Modes
{{-- Interactive (default) - user can rotate/zoom --}}
<livewire:molecule smiles="CCO" mode="interactive" />
{{-- Rotating - auto-rotates on Y axis --}}
<livewire:molecule smiles="CCO" mode="rotating" />
{{-- Static - no interaction --}}
<livewire:molecule smiles="CCO" mode="static" />
Visualization Styles
<livewire:molecule smiles="CCO" style="stick" />
<livewire:molecule smiles="CCO" style="sphere" />
<livewire:molecule smiles="CCO" style="ball-and-stick" />
<livewire:molecule smiles="CCO" style="cartoon" /> {{-- Best for proteins --}}
<livewire:molecule smiles="CCO" style="line" />
Customizing Appearance
<livewire:molecule
smiles="c1ccccc1"
mode="interactive"
style="ball-and-stick"
background-color="#1a1a2e"
width="500px"
height="400px"
/>
Advanced 3Dmol Options
Pass additional 3Dmol.js options through the Livewire wrapper:
<livewire:molecule
smiles="CCO"
:viewer-options="['backgroundAlpha' => 0.0, 'disableFog' => true]"
:model-options="['keepH' => true]"
:style-options="['stick' => ['radius' => 0.2]]"
/>
Reactive Updates
The component reacts to property changes:
<div x-data="{ currentSmiles: 'CCO' }">
<select x-model="currentSmiles">
<option value="CCO">Ethanol</option>
<option value="CC(=O)O">Acetic Acid</option>
<option value="c1ccccc1">Benzene</option>
</select>
<livewire:molecule :smiles="$currentSmiles" />
</div>
Configuration
// config/livewire-molecule.php
return [
// Default background color
'default_background' => '#ffffff',
// HTTP timeout for external APIs (seconds)
'timeout' => 10,
// Default 3Dmol.js options
'viewer_options' => [],
'model_options' => [],
'style_options' => [],
// Cache settings for resolved molecules
'cache' => [
'enabled' => true,
'ttl' => 60 * 60 * 24, // 24 hours
'prefix' => 'molecule_',
],
];
Input Format Priority
When multiple identifiers are provided, the component uses this priority:
sdf(raw data, no API call needed)pdb(fetches from RCSB PDB)pubchem-cid(fetches from PubChem)smiles(converts via NCI CACTUS)inchi(converts via NCI CACTUS)
External APIs Used
This package relies on these free public APIs for structure retrieval and conversion:
| API | Purpose | Rate Limits |
|---|---|---|
| RCSB PDB | Fetch protein structures | Generous |
| PubChem | Fetch compound structures | 5 req/sec |
| NCI CACTUS | SMILES/InChI → 3D conversion | Best effort |
For production use with high traffic, consider implementing your own conversion service or caching aggressively.
Troubleshooting
"Failed to convert SMILES to 3D structure"
- Verify the SMILES string is valid
- The NCI CACTUS service may be temporarily unavailable
- Some complex molecules may fail to convert
Molecule appears blank
- Check browser console for JavaScript errors
- Ensure 3DMol.js is loading (check Network tab)
- Verify the molecule data is being resolved (check
$moleculeDataproperty)
"Cannot connect to [API]" or blank molecule in production
This package requires outbound HTTP access to external APIs for SMILES/InChI conversion and PubChem/PDB data fetching.
Laravel Vapor: Enable outbound HTTP in your vapor.yml:
id: your-project-id
name: your-project-name
environments:
production:
egress: true # Enable outbound HTTP
Other platforms: Ensure your server/firewall allows outbound HTTPS to:
cactus.nci.nih.gov(SMILES/InChI conversion)pubchem.ncbi.nlm.nih.gov(PubChem data)files.rcsb.org(PDB structures)3dmol.csb.pitt.edu(3DMol.js CDN)
Workaround: Use raw SDF/PDB data instead of SMILES/PubChem CIDs to avoid external API calls:
<livewire:molecule :sdf="$sdfData" />
Upgrade Guide (v1 → v2)
v2 renames the publishable config file and config key.
- Republish config:
php artisan vendor:publish --tag=molecule-config
- Update config usage:
- File moved from
config/molecule.phptoconfig/livewire-molecule.php - Config key changed from
moleculetolivewire-molecule
If you referenced config values in your app, update:
// v1
config('molecule.default_background');
// v2
config('livewire-molecule.default_background');
Local Development
To preview the component in a real browser while working on the package:
composer serve
Then open http://localhost:8000 to see a live demo page with multiple molecule examples rendered using the actual component.
Testing
composer test
Acknowledgments
- 3DMol.js - BSD-3-Clause licensed molecular viewer
- NCI CACTUS - Chemical structure conversion service
- PubChem - Chemical compound database
License
MIT License. See LICENSE for details.
This package includes 3DMol.js which is licensed under BSD-3-Clause.